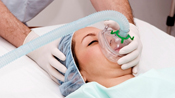
Smith senior Germaine Nendah aims to identify genes in E. coli whose expression changes at fever temperature allow the organisms to survive.
/ Published February 11, 2011
Many Smith seniors studying in the sciences are using their final year to dig deep,
ask questions, and conduct original research for honors theses that open new horizons and
make discoveries new to science. Insight has profiled three of those projects.
Although fever is one of the body’s best defenses against invading pathogens, many bacteria have learned how to stand the heat. Organisms such as Escherichia coli can survive a fever by increasing the expression of certain genes and thereby amplifying proteins that improve their chances of survival. Understanding exactly which genes allow pathogens to evade the immune system’s fever response may lead to the development of therapeutic drugs that complement the body’s own defenses.

Germaine Nendah ’11
“If we know what the bacteria are doing to continue infecting the host, then we know how to design drugs to help the immune response,” explains Germaine Nendah ’11. Those drugs would target only the proteins that allow for the expression of certain vital genes instead of wasting resources on those proteins that are not essential for survival when bacteria are attacked by the immune system.
Working on her honors thesis in the microbiology lab of Professor Christine White-Ziegler, Nendah aims to identify genes in E. coli whose expression changes at fever temperatu re allow the organisms to survive. She is now testing two subsets of genes typically associated with virulence in bacteria that E. coli use to efficiently colonize their human host. Because pathogenic strains of E. coli are among the primary causes of urinary tract infections in women and the largest cause of nosocomial infections in the United States, successfully fighting infections caused by these bacteria is especially critical, Nendah says.
When the immune system is alerted of a bacterial infection, body temperature increases to both slow down and kill the invading pathogens. Higher temperatures limit the minerals available in the circulating blood stream, which bacteria require to grow and divide. Fever conditions also damage bacterial cell walls, and this causes an imbalance between the inside and outside of the cell, leading to cell death.
Nendah thinks that in response to harsh fever conditions, bacteria may increase the expression of certain genes that allow the bacteria to continue infecting their human hosts. While the bacteria may not exactly thrive, she says, the fact that 10 percent of the E. coli genome, a total of 432 genes, are temperature-regulated, suggests that these pathogens are able to survive by increasing or decreasing the expression of some of their genes in response to a fever.
The most familiar players in evading the destructive effects of temperature change are heat-shock proteins, which, as their name suggests, allow bacteria to cope with extreme conditions by keeping other vital proteins intact. Nendah is determining the effects of a fever temperature on the expression of two other groups of genes, called fimbria (pap) and ion-acquisition genes (fes, fhuA, and cirA). The pap gene is responsible for producing an adhesin protein, which allows the bacteria to more tightly grip the epithelial cell lining of the urinary tract. Iron-acquisition-genes increase the iron uptake that is essential for cell metabolism and prevents slowing of motility and replication brought on by fever.
Nendah grows strains of bacteria at 37°C (99°F), normal human body temperature, and then shifts them to 40°C (104°F) to mimic a high fever. By isolating all the RNA from these organisms at 30-minute intervals, she monitors the expression of specific genes over an eight-hour period.
Nendah’s initial observations on expression levels of iron genes—from a nonpathogenic strain of E. coli that does not harm humans—have surprised her, she explains. Expression of the pap gene did not change with a 3°C increase in temperature, suggesting that expression of this gene may not be affected by a fever. While ion-acquisition genes were up-regulated at fever temperature, a complex pattern of increase and decrease in expression over eight hours revealed that other factors, such as iron levels in the bacteria, may also be responsible for regulating expression of this group of genes.
In January, Nendah will shift her studies from a comensal strain of E. coli, which does not infect humans, to a pathogenic one and may extend her search to other genes that produce proteins involved in motility and heat-shock functions.
The more up-regulated genes that can be identified, the more chances scientists have of designing a drug powerful enough to wipe out invading pathogens capable of evading the body’s normal immune response, Nendah explains. “We don’t need to spend millions of dollars to design antibiotics that bacteria become resistant to in the long-run. We need to design drugs that will completely prevent the ability of bacteria to produce proteins from specific genes essential for survival despite immune defenses.”